
Here, we discuss the molecular description and phylogenetic analysis of a potential third pathogenic Orientia species detected in 18 patients with scrub typhus in southern Chile. Genomic information on Orientia strains from Chile has been insufficient and scrub typhus in the Middle East region is caused by a new Orientia species, Candidatus Orientia chuto ( 6), highlighting that our current knowledge on the spectrum of Orientia species is incomplete ( 13). This paradigm, however, has recently been brought into question with evidence of scrub typhus being found in the Middle East, Africa, and South America ( 7– 12). Disinterest has been influenced by the perception that scrub typhus is a geographically limited disease, threatening rural populations within a certain region, known as the “tsutsugamushi triangle,” and rarely affecting travelers ( 5, 6). Although this disease has been known since at least 313 ce and currently threatens over 1 billion people in Asia and Australasia, it is widely underdiagnosed and underreported ( 3, 4). Scrub typhus is caused by Orientia tsutsugamushi, a strictly intracellular bacterium with a remarkable genetic and antigenic diversity ( 1, 2). Scrub typhus is a potentially fatal rickettsial infection transmitted by larval stage trombiculid mites called chiggers. Our results indicate that Orientia isolates from Chile constitute a novel species, which, until they are cultivated and fully characterized, we propose to designate as Candidatus Orientia chiloensis, after the Chiloé Archipelago where the pathogen was identified. Phylogenetic analyses of both genes grouped the specimens from Chile in a different clade from other Orientia species. tsusugamushi and 3.0% for rrs and 14.8% for htrA compared with Candidatus Orientia chuto. Their diversity was 3.1%–3.5% for rrs and 11.2%–11.8% for htrA compared with O. Sequences were ≥99.7% identical among the samples for both amplified genes. We analyzed Orientia gene sequences of 16S rRNA ( rrs) and 47-kDa ( htrA) from 18 scrub typhus patients from Chile. Although considered to be restricted to the Asia Pacific region, scrub typhus has recently been discovered in southern Chile. Staphylococcus aureus coagulase restriction fragment length polymorphism sequence-based phylogenetic analysis.Scrub typhus is a potentially fatal rickettsiosis caused by Orientia species intracellular bacteria of the genus Orientia. The coa genotypes and their restriction patterns observed in the present study are novel, not published earlier.
Two coa patterns were observed in mastitic milk indicating multiple origins of infection, with 595 bp coa genotype being predominant in mastitic milk. aureus coagulase typeshave a site-specific predilection. The study, being localized to only one farm, yielded different RFLP patterns as observed from different sampling sites, which indicates that different S. While as the most distant sequences with the value of 0.483 were found between S.
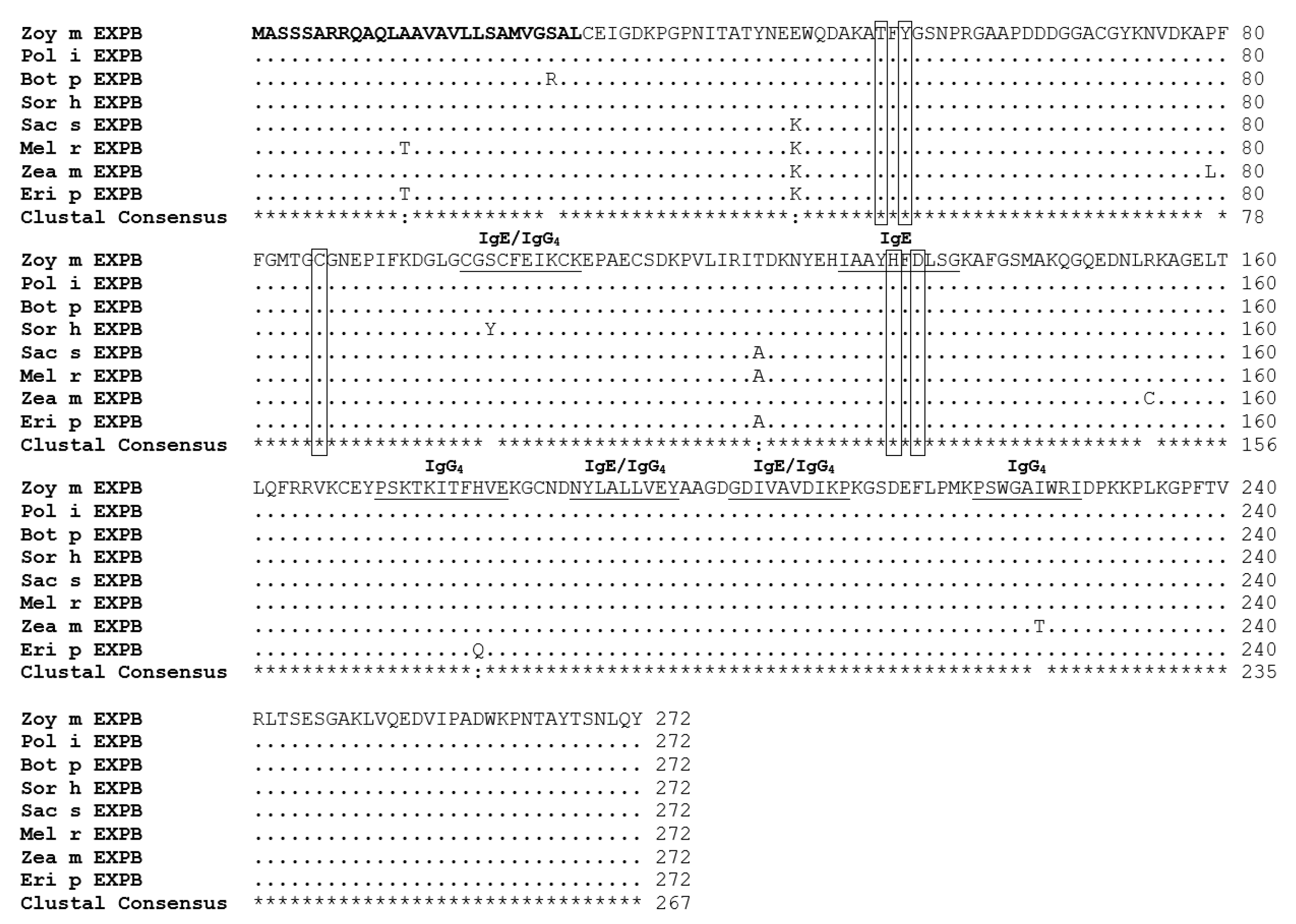
aureus 595 (mastitic milk), and thus, they are most closely related. The most identical sequences with the value of 0.810 were found between isolate S. On sequencing, four different sequences having unique restriction patterns were obtained. On RFLP using both restriction enzymes, four different restriction patterns P1, P2, P3, and P4 were observed.
#Bioedit sequence identity matrix skin#
514 bp (2 isolates) and 757 bp (4 isolates) coa genotypes were observed from nasal cavity and pus from skin wounds, respectively. Two coa genotypes, 595 bp (15 isolates) and 802 bp (4 isolates), were observed in mastitic milk. Four different genotypes of coa gene, i.e., 514 bp, 595 bp, 757 bp, and 802 bp were obtained. aureus isolates, 25 (64.10%) isolates carried coa gene. aureus were confirmed by targeting nuc gene using PCR. Of 192 different samples,39 (20.31%) isolates of S. These sequences were aligned for maximum homology using the Bioedit softwareandsimilarity in the sequences was inferred with the help of sequence identity matrix. One isolate from each restriction pattern was sequenced. Different coa genotypes observed were subjected to RFLP using restriction enzymes Hae111 and Alu1, to obtain the different restriction patterns. aureus isolates were subjected to coagulase ( coa) gene PCR. The presumptive isolates were confirmed by nuc gene-based polymerase chain reaction (PCR). This study was conducted to study the coagulase gene-based genetic diversity of Staphylococcus aureus, isolated from different samples of cattle using restriction fragment length polymorphism (RFLP) and their sequence-based phylogenetic analysis.Ī total of 192 different samples from mastitic milk, nasal cavity, and pus from skin wounds of cattle from Military Dairy Farm, Jammu, India, were screened for the presence of S.
